Updated 9 November 2018 with corrected use of the left-censored survival model
This is a post about dealing with tables that have been subject to cell suppression or perturbation for confidentiality reasons. While some of the data world is only just waking up to issues of confidentiality and privacy, national statistics offices and associated researchers over some decades have developed a whole field of knowledge and tools in the area of “statistical disclosure control”. I found Duncan, Elliot and Salazar-Gonzalez’ Statistical Confidentiality: Principles and Practice and excellent text on this subject when I had to get on top of it late in 2017, while working on Stats NZ’s Integrated Data Infrastructure (IDI) output release processes.
Why does one need to suppress data when it has already been aggregated and isn’t sensitive itself? Because we fear “data snoopers” (yes, that really is the correct technical term) who will combine our published data with other datasets - public or private - to reveal sensitive information. For example, if we publish a table that shows there is only one hairdresser in Smallville, that in itself isn’t sensitive; but if we combine it with a separate dataset showing the average income of hairdressers in Smallville we have given away their private information. The thing that makes statistical disclosure control so hard is that we don’t know what other datasets the data snooper may have available to them to use in combination with our data, however innocuous it looks on its own.
Suppression in context
National stats offices rarely rely alone on cell suppression. One reason is that unless analysis is very very tightly controlled, the secondary suppressions that are needed to avoid uncovering the true value of a suppressed cell quickly get out of hand. For example, in this table which I’ll be returning to later, we would have to suppress not only number of tigers and bears in region B, but also the total number of tigers and bears:
| region | lions | tigers | bears |
|---|---|---|---|
| A | 7 | 6 | 13 |
| B | 9 | <6 | <6 |
| C | 20 | 24 | 33 |
| D | 18 | 8 | 90 |
If it were critical that we publish the total number of tigers and bears, then we would need to suppress at least one more region count for each animal. So the numbers of tigers and bears in Region A would probably also be suppressed, not because it is less than six but because revealing it would let a snooper with the marginal totals deduce the counts in Region B. With the hierarchical tables and various slices and dices that official statistics practice usually requires, this procedure soon gets pretty complicated.
There is software to help calculate the best secondary cell suppressions (such as the sdcTable R package), but sometimes there are practical situations in which it would be difficult to apply, and for this or other reasons we need more techniques than just suppression. One key situation is when a significant number of analysts, not under the direct control of the agency, have a degree of access to the original microdata. Two examples I’m aware of are:
- the Australian Bureau of Statistics’ Census Table Builder is an online tool (with free guest access) that lets you slice and dice granular census microdata, without seeing the original data but delivering aggregate tables of counts. The ABS can’t control which combinations of filters and slices will be applied, so secondary cell suppression isn’t realistic. Instead, results are reported all the way down to zero, but have been perturbed so one never knows exactly what the correct value is (other than within a small margin, of course).
- the Stats NZ data lab allows access to de-identified microdata including the extraordinary Integrated Data Infrastructure which combines inter alia tax, benefits, health, education and justice data at the individual level. While the data are rigorously de-identified (researchers cannot see names, tax or other identification numbers, or addresses more granular than meshblock), a snooper with full access could probably combine results from the IDI with external datasets to identify sensitive information. Because of this, access is tightly controlled to secure laboratories and approved researchers, who are responsible for confidentialisation of their results. Results have to go through an “output checking” process to be sure appropriate confidentialisation has been applied. There is a 37 page Microdata Output Guide for researchers to follow throughout this process. Many of the rules involve suppression and secondary suppression, but these are not the only tools relied on. For example, counts are perturbed by means of random rounding.
Random rounding is an interesting case. Base 3 random rounding works by rounding a number to the nearest number divisible by 3 with 2/3 probability, and the second nearest with 1/3. If the number is already divisible by 3 it stays the same. So this means that of all the numbers that might end up being published as a six, “4” has a 1/3 chance of being 6; “5” has 2/3; “6” has 3/3; “7” has 2/3; and “8” has 1/3.
Simple application of Bayes’ formula indicates that if we started with equal chances for the real value being anything from 4 to 8, then the probabilities of a published six really meaning 4, 5, 6, 7 and 8 are 1/9, 2/9, 3/9, 2/9, 1/9 respectively. But were the starting chances equal? Not really. In fact, I’m sure the number “8” occurs in the sort of cross tabs that get cell suppression many times more than the number “4”. I don’t have proof of this, just an intuition based on the process of deciding what categories are used in the first place, which tables are thought worth publishing, etc. Obviously if half the numbers in a table were small enough to be suppressed, pretty soon we would stop bothering to publish the table and find other categories with higher cell counts.
If we change our priors from equal chances to something more realistic, we get quite a different picture:
The inference process here can be thought of as: “Before we even see the table, we know it is a table of counts. We know that most published counts of this sort are substantial numbers, at least in double figures. Then we look at this particular cell, and see that it is published as a random rounded 6. We update our prior, with all possibilities lower than 4 and greater than 8 automatically becoming zero. Then we combine our updated prior with the chances of each value being published as 6, and get a posterior distribution for the underlying number that relies on both our starting assumptions and the process of random rounding.”
We’ll come back to this line of thinking when we impute some numbers that are suppressed completely.
Here’s the code that generated the graphic above and sets us up for the rest of the session.
library(tidyverse)
library(scales)
library(knitr)
library(MASS) # for corresp
library(Cairo)
library(png)
library(VGAM)
library(grid)
library(ggrepel)
library(mice)
library(testthat)
#---------random rounding base 3-----------------
# probability of observing a six, given actual values 4, 5, 6, 7, 8
prob <- c(1,2,3,2,1) / 3
prior15 <- dpois(4:8, 15)
prior30 <- dpois(4:8, 30)
rr3_data <- data_frame(
Value = 4:8,
`Equal probabilities` = prob / sum(prob),
`Poisson with mean of 15` = prob * prior15 / sum(prior15 * prob),
`Poisson with mean of 30` = prob * prior30 / sum(prior30 * prob)
) %>%
gather(prior, post_prob, -Value)
# check our probabilities add up to 1
expect_equal(as.vector(tapply(rr3_data$post_prob, rr3_data$prior, sum)),
c(1,1,1))
rr3_data %>%
ggplot(aes(x = Value, weight = post_prob)) +
geom_bar(fill = "steelblue", alpha = 0.8) +
facet_wrap(~prior) +
labs(y = "Posterior probability this is the actual value") +
ggtitle("A value that has been random rounded (base 3) for statistical disclosure control",
"Distribution of true value when published value is 6, with three different prior expectations")Simulated suppressed data
Even though national stats offices rarely rely just on cell suppression, agencies that need to safely publish aggregate data but lack the resources and expertise of a stats office will often use it as their main method to provide a certain minimal level of confidentialisation. So it’s of some interest to look at the case of how to analyse a table that has had cells suppressed but no other statistical disclosure control. I am using two such datasets, from completely different sources on different projects, in my work at the moment.
Earlier in this post I introduced this table with two suppressed cells:
Data as it is published
| region | lions | tigers | bears |
|---|---|---|---|
| A | 7 | 6 | 13 |
| B | 9 | <6 | <6 |
| C | 20 | 24 | 33 |
| D | 18 | 8 | 90 |
I generated this from a model with a strong relationship between animal type and region, and I also know both the true observed values (as though I were the statistical office managing suppression and publication) and the expected values from the data generating process (as though I were God).
The actual data
| region | lions | tigers | bears |
|---|---|---|---|
| A | 7 | 6 | 13 |
| B | 9 | 1 | 5 |
| C | 20 | 24 | 33 |
| D | 18 | 8 | 90 |
The expected values from the data generating process
| region | lions | tigers | bears |
|---|---|---|---|
| A | 7.4 | 4.8 | 13.2 |
| B | 6.2 | 1.6 | 11.6 |
| C | 15.9 | 22.8 | 31.4 |
| D | 17.8 | 10.7 | 79.1 |
The data was generated by means of a generalized linear model with a Poisson response (a classic and critical model to understand when dealing with tables of count data), this way:
data <- expand.grid(animals = c("lions", "tigers", "bears"),
region = LETTERS[1:4],
count = 1) %>%
as_tibble() %>%
mutate(animals = fct_relevel(animals, "lions"))
n <- nrow(data)
mm <- model.matrix(count ~ animals * region, data = data)
# for reproducibility:
set.seed(123)
# generate true coefficients of the model:
true_coefs <- c(2, runif(n - 1, -1, 1))
# generate the actual data and make the suppressed version:
data <- data %>%
mutate(expected = exp(mm %*% true_coefs),
count = rpois(n, lambda = exp(mm %*% true_coefs)),
censored_count = ifelse(count < 6, "<6", count),
censored_count_num = as.numeric(censored_count),
count_replaced = ifelse(count < 6, 3, count))
# data as it is published:
data %>%
dplyr::select(animals, region, censored_count) %>%
spread(animals, censored_count) %>%
kable
# actual underlying data:
data %>%
dplyr::select(animals, region, count) %>%
spread(animals, count) %>%
kable
# actual underlying data generating process expectations:
data %>%
dplyr::select(animals, region, expected) %>%
mutate(expected = round(expected, 1)) %>%
spread(animals, expected) %>%
kableCorrespondence analysis and tests of independence
The most common statistical test done on this sort of data is a Chi-square test, which tests the plausibility of the row and column variables being independent of each other. It’s equivalent to comparing a generalized linear model with a Poisson response that has an interaction effect between two variables (count ~ animals + region + animals:region) with one that doesn’t (count ~ animals + region). As I generated the data with the fully saturated model, I know that the correct decision is to reject the null hypothesis of “no interaction”. But what’s a general way to perform this test on a table with data that is missing, very much not at random but missing because it is below a known threshold?
A second technique I want to look at is correspondence analysis, coming from a more exploratory paradigm of data analysis rather than basing itself on statistical tests. Correspondence analysis works well as a graphic summary of a two-way cross tab. Here’s the biplot from correspondence analysis of the true observed data in our case:
If you’re not familiar with this sort of plot and you deal with count data in cross tabs, I strongly recommend the technique. At a glance we see that bears are distinctively frequent to area D, tigers to C and lions B and A (although A is a more typical region than others, close to the cross in the centre of the chart). You can confirm this by looking at the original table and calculating row-wise and column-wise percentages, but it’s not such an easy snapshot as is provided by the graphic.
Like the Chi-square test, we need a full table of results to do this analysis.
With only two cells suppressed, there are only 36 possibilities for what they could be (each cell could really be any number from zero to five inclusive). So it’s quite feasible to perform analysis on 36 different possible data sets. If the results are substantively the same for all combinations, we can conclude it doesn’t matter what the suppressed cells really are. This is a nice, common sense and easily explainable approach.
For the correspondence analysis, the logical way to show all 36 possibilities is via an animation:
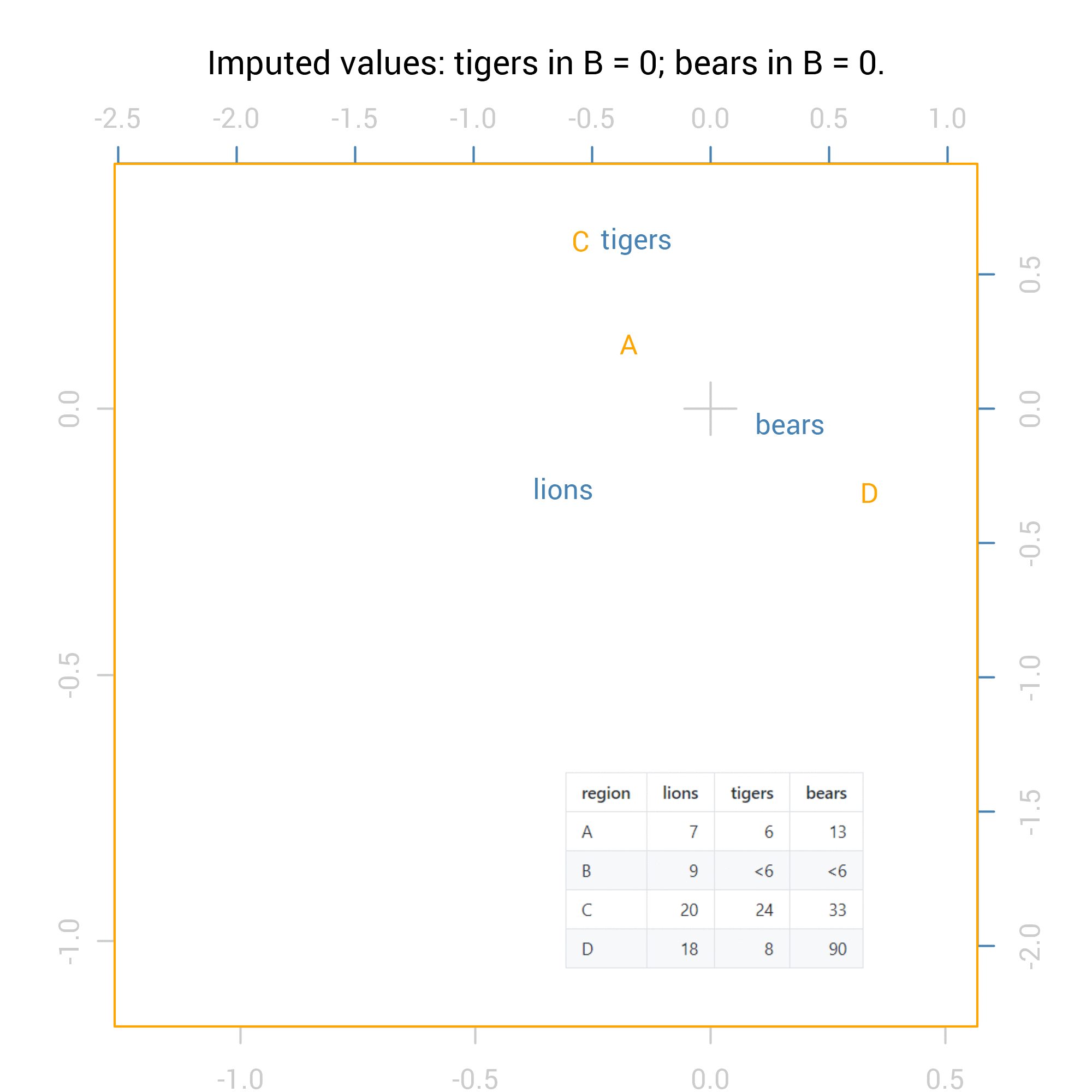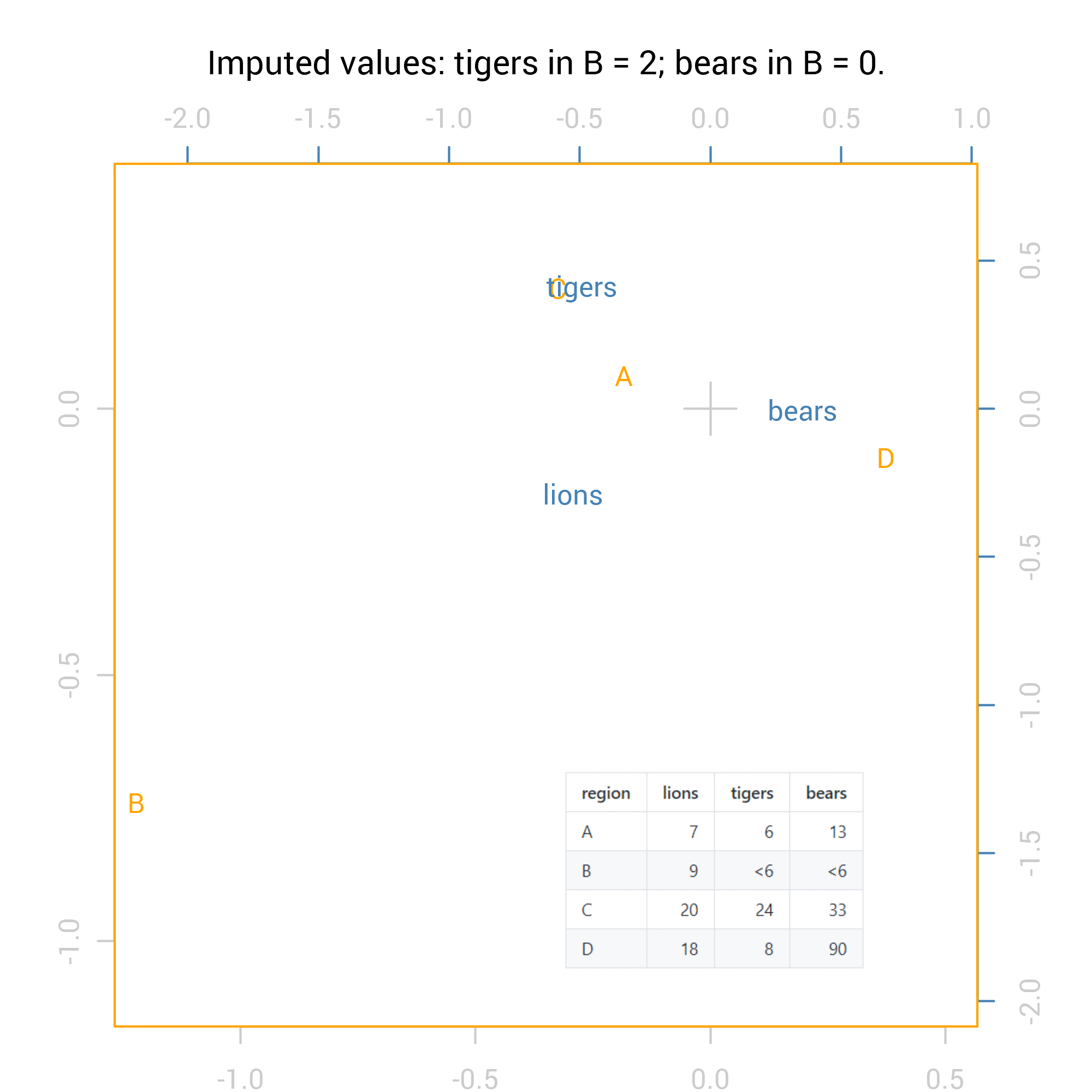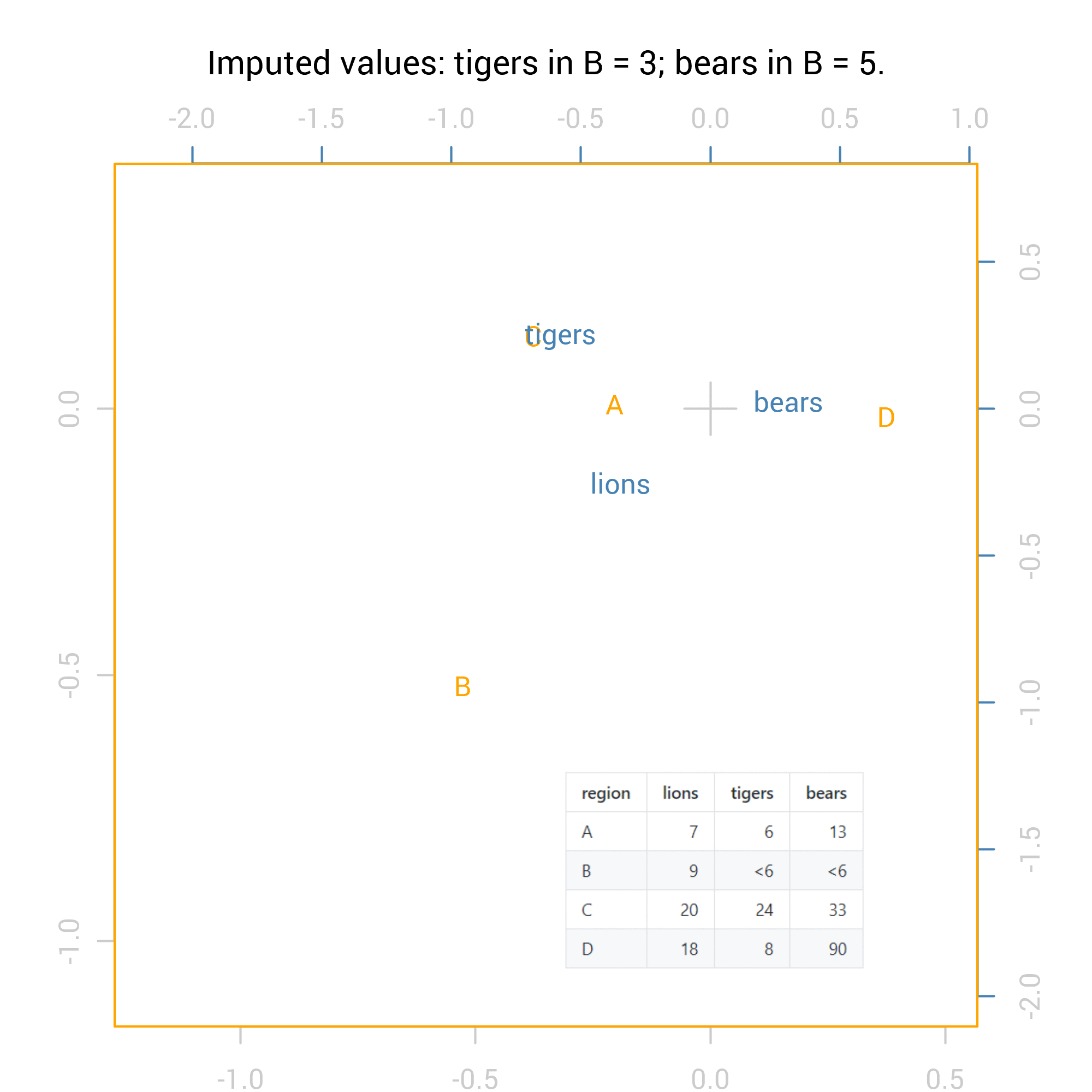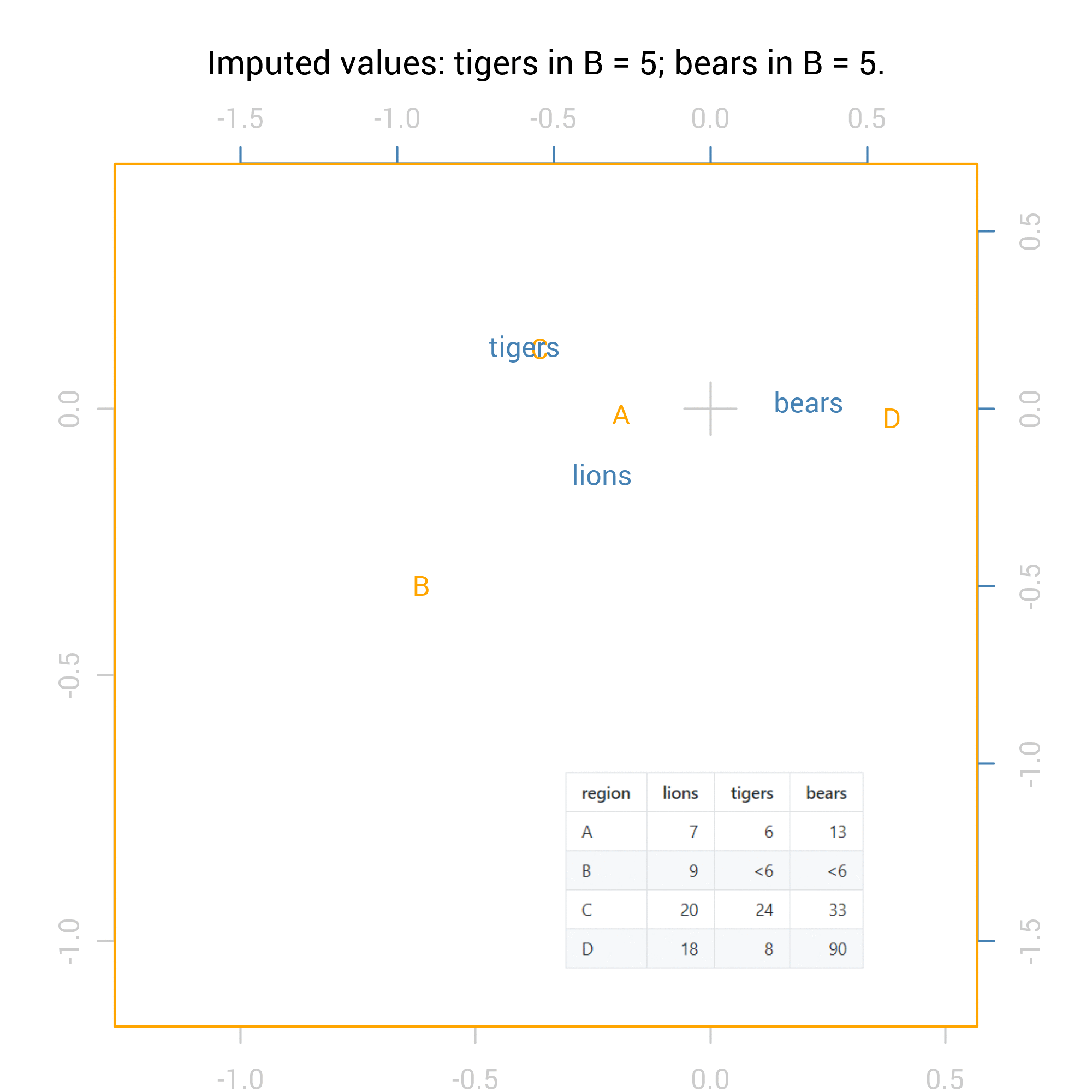
For the Chi square tests the results are numeric so we can summarise them with standard methods including maximum and minimum values. Here are the summaries of the p values from the Fisher exact test for independence (pvalues_f) and the Chi-squared test:
pvalues_f pvalues_c
Min. :0.0004998 Min. :4.810e-11
1st Qu.:0.0004998 1st Qu.:6.381e-09
Median :0.0004998 Median :3.989e-08
Mean :0.0004998 Mean :1.480e-07
3rd Qu.:0.0004998 3rd Qu.:1.730e-07
Max. :0.0004998 Max. :8.438e-07
For both the correspondence analysis and the tests of independence, the results don’t vary materially when we change the values in the suppressed cells, so we could proceed to inference with confidence.
Here’s the code that did those two types of analysis with a full set of possible values plugged into the suppressed cells:
# make a traditional cross tab matrix of the data:
tab <- data %>%
dplyr::select(animals, region, count) %>%
spread(animals, count) %>%
dplyr::select(lions, tigers, bears) %>%
as.matrix()
rownames(tab) <- LETTERS[1:4]
# biplot with the actual full values (obviously not usually available to analysts):
par(font.main = 1, col.axis = "grey80", fg = "grey80")
xl <- c(-1.2, 0.5)
yl <- c(-1.1, 0.4)
biplot(corresp(tab, nf = 2),
main = "Full data including suppressed values\n",
col = c("orange", "steelblue"),,
xlim = xl, ylim = yl)
# read in image of the table for including in the animated graphic:
table_image <- readPNG("0138-table-image.png")
# set up empty vectors to store the results of tests of independence:
pvalues_f <- numeric()
pvalues_c <- numeric()
# create a temporary folder for storing frames of the animation:
dir.create("tmp")
pd <- setwd("tmp")
# we can use brute force and see if the results are statistically significant
# for all possible values. We'll do chi square tests and correspondence analysis
# at the same time
for(i in 0:5){
for(j in 0:5){
tab[2, 2] <- i # tigers in B
tab[2, 3] <- j # bears in B
pvalues_f <- c(pvalues_f, fisher.test(tab, simulate.p.value = TRUE)$p.value)
pvalues_c <- c(pvalues_c, chisq.test(tab)$p.value)
CairoPNG(paste0(i, j, ".png"), 2000, 2000, res = 300)
par(font.main = 1, col.axis = "grey80", fg = "grey80", mar = c(2,1,4,1))
biplot(corresp(tab, nf = 2), col = c("orange", "steelblue"),
main = paste0("Imputed values: tigers in B = ", i, "; bears in B = ", j, ".\n"),
xlim = xl, bty = "n",
ylim = yl)
grid.raster(table_image, x = 0.66, y = 0.2, width = 0.3)
dev.off()
}
}
system('magick -loop 0 -delay 40 *.png "0138-corresp.gif"')
# move the result to where I use it for the blog
file.rename("0138-corresp.gif", "../../img/0138-corresp.gif")
setwd(pd)
# clean up (uncomment the last line if happy to delete the above results):
# unlink("tmp", recursive = TRUE)
# p-values:
summary(cbind(pvalues_f, pvalues_c))Generalized linear model
For a more general approach, I wanted to be ready for a situation with more than two categorical variables, and where the number of suppressed cells is too large to repeat the analysis with all combinations. As a use case, I fit a generalized linear model where my interest is in the values of the coefficients (ie the tiger effect relative to lions, bear effect, region B effect relative to region A, etc and their interaction impacts, on “count” as the response variable).
One thought is to treat the missing data as a parameter, constrained to be an integer from zero to five, and estimate it simultaneously with the main model in a Bayesian framework. In principle this would work, except the missing data has to be an integer (because it then forms part of a Poisson or negative binomial distribution as part of the main model) and it is difficult to estimate such discontinuous parameters. You can’t do it in Stan, for example.
Another way of looking at the problem is to note that this is censored data and use methods developed specifically with this in mind. “Survival analysis” has of course developed methods for dealing with all sorts of censored data; most obviously with “right-censored” data such as age at death when some of the subjects are still alive. Left-censored data occurs when measurement instruments struggle at the lower end of values, but is also an appropriate description of our suppressed values. In R, the VGAM package provides methods for fitting generalized linear models to data that is either left, right or interval censored.
The third family of methods I tried involved substituting values for the suppressed cells. At its simplest, I tried replacing them all with “3”. At the more complex end, I used multiple imputation to generate 20 different datasets with random values from zero to five in each suppressed cell; fit the model to each dataset; and pool the results. This is a method I use a lot with other missing data and is nicely supported by the mice R package. mice doesn’t have an imputation method (that I can see) that exactly meets my case of a number from zero to five, but it’s easy to write your own. I did this two different ways:
- with a uniform chance for each number
- with probabilities proportional to the density of a Poisson distribution with mean equal to the trimmed mean of the unsuppressed counts (the reasoning being the same as that used earlier in this post in discussing random rounding)
Here’s the results of all those different methods. First, in a summary of the mean squared error comparing coefficient estimates to their known true values:
… and second, with the coefficient-by-coefficient detail:
What we see from this single simulated dataset is:
- simply substituting the number “3” in for all suppressed values worked fine; in fact nearly as well as using the original unavailable data, or the most sophisticated multiple imputation method
- multiple imputation with probabilities proportional to the Poisson density out-performed multiple imputation with sampling from a uniform distribution.
- the survival analysis method performed ok by this crude measure (in fact, better than fitting regression to the full data), once I got my indicator of which observations were censored the right way around, thanks to the comment by Aniko Szabo - but not as well as the best multiple imputation method.
- Sampling the suppressed values from a uniform distribution, and the left-censored survival analysis method, both give particularly bad coefficients for the coefficients related to the suppressed cells (interaction of tigers with region B, and bears with region B).
Because this is a single simulated dataset, the conclusions above should be treated as only indicative. A good next step would be to generalise this by testing it with many datasets, but that will have to wait for a later blog post as this one is already long enough! My working hypothesis when I get to that is that multiple imputation with probabilities proportional to the Poisson density will be the best method, but I’m open to being wrong on that.
Here’s the code for the comparison of those different methods:
#====================different approaches to dealing with the missing data===================
#---------------regenerate data-----------
# doing this here to make this chunk of code easier for when I want to generalise this whole approach
# to multiple simulations
set.seed(123)
true_coefs <- c(2, runif(n - 1, -1, 1))
data <- data %>%
mutate(expected = exp(mm %*% true_coefs),
count = rpois(n, lambda = exp(mm %*% true_coefs)),
censored_count = ifelse(count < 6, "<6", count),
censored_count_num = as.numeric(censored_count),
count_replaced = ifelse(count < 6, 3, count))
#--------------straightforward methods-----------
mod0 <- glm(count ~ animals * region, data = data, family = "poisson")
mod2 <- glm(count_replaced ~ animals * region, data = data, family = "poisson")
#------------with censored poisson regression-------------------------
# (note that we indicated a complete observation with a 1, and a censored one with a 0)
data$z <- pmax(6, data$count)
data$not_lcensored <- as.numeric(!is.na(data$censored_count_num ))
mod3 <- vglm(SurvS4(z, not_lcensored, type = "left") ~ animals * region, family = cens.poisson, data = data)
#------------------ with multiple imputation----------
#' Imputation function for suppressed data for use with mice - Poisson-based
#'
#' @param y vector to be imputed
#' @param ry logical vector of observed (TRUE) and missing (FALSE) values of y
#' @param x design matrix. Ignored but is probably needed for mice to use.
#' @param wy vector that is TRUE where imputations are created for y. Not sure when this is different
#' to just !ry (which is the default).
mice.impute.desuppress <- function (y, ry, x, wy = NULL, max_value = 5, ...) {
# during dev:
# y <- data$censored_count_num; ry <- !is.na(y)
if (is.null(wy)){
wy <- !ry
}
# What are the relative chances of getting values from 0 to the maximum allowed value,
# if we have a Poisson distribution which we have estimated the mean of via trimmed mean?
# (this is very approximate but probably better than just giving equal chances to 0:5)
probs <- dpois(0:max_value, mean(y[ry], tr = 0.2))
return(sample(0:max_value, sum(wy), prob = probs, replace = TRUE))
}
#' Imputation function for suppressed data for use with mice - simple
#'
mice.impute.uniform <- function (y, ry, x, wy = NULL, max_value = 5, ...) {
return(sample(0:max_value, sum(wy), replace = TRUE))
}
m <- 20 # number of imputed/complete datasets to generate
data_imp1 <- mice(data[ , c("censored_count_num", "animals", "region")],
method = "desuppress", print = FALSE, m = m)
imp_y1 <- data_imp1$imp$`censored_count_num`
imp_y1 # visual check
data_imp2 <- mice(data[ , c("censored_count_num", "animals", "region")],
method = "uniform", print = FALSE, m = m)
imp_y2 <- data_imp2$imp$`censored_count_num`
imp_y2 # visual check
mod_mice1 <- with(data_imp1, glm(censored_count_num ~ animals * region, family = "poisson"))
coef_mice1 <- pool(mod_mice1)$pooled$estimate
mod_mice2 <- with(data_imp2, glm(censored_count_num ~ animals * region, family = "poisson"))
coef_mice2 <- pool(mod_mice2)$pooled$estimate
#---------------results----------------------
# comparison data
d <- data_frame(underlying = true_coefs,
`Using full data including suppressed values (not usually possible!)` = coef(mod0),
`Replacing suppressed values with 3` = coef(mod2),
`Left-censored survival-based method` = coef(mod3),
`MICE with Poisson proportional probabilities` = coef_mice1,
`MICE with uniform probabilities` = coef_mice2,
labels = names(coef(mod0))) %>%
mutate(labels = gsub("animals", "", labels),
labels = gsub("region", "", labels)) %>%
gather(method, value, -underlying, -labels) %>%
mutate(method = str_wrap(method, 25)) %>%
mutate(value = ifelse(is.na(value), 0, value))
# summary data:
d2 <- d %>%
mutate(square_error = (value - underlying) ^ 2) %>%
group_by(method) %>%
summarise(mse = mean(square_error),
trmse = mean(square_error, tr = 0.2)) %>%
ungroup() %>%
mutate(method = fct_reorder(method, mse))
# summary graphic:
ggplot(d2, aes(x = mse, y = trmse, label = method)) +
geom_point(size = 2) +
geom_text_repel(colour = "steelblue") +
labs(x = "Mean squared error of coefficient estimates",
y = "Trimmed mean squared error of coefficient estimates") +
ggtitle("Comparison of different methods of handling suppressed counts in a frequency table",
"Summary results of models fitted after different treatment methods")
# coefficients graphic:
d %>%
mutate(method = factor(method, levels = levels(d2$method))) %>%
ggplot(aes(x = underlying, y = value, label = labels)) +
facet_wrap(~method, nrow = 1) +
geom_abline(slope = 1, intercept = 0, colour = "grey50") +
geom_point() +
geom_text_repel(colour = "steelblue") +
labs(x = "Correct value of coefficient in underlying data generating process",
y = "Value estimated from a GLM with one particular method of dealing with censored data") +
coord_equal() +
ggtitle("Comparison of different methods of handling suppressed counts in a frequency table",
"Coefficient estimates from models fitted after different treatment methods")


